Project Overview
This project explores how different image modalities influence the performance of machine learning models when analyzing Chironomus alluaudi, a forensically important non-biting midge used in post-mortem interval (PMI) estimation. Identifying larval stages of this species is challenging because it often relies on expert taxonomic knowledge and subtle morphological features.
Using a convolutional autoencoder (CAE), this study compares three image modalities—color, grayscale, and negative—to determine how they affect feature extraction and model performance. The analysis evaluates clustering of latent features, reconstruction error patterns, and edge detection metrics.
Results show that image modality significantly influences machine learning performance. The negative image modality produced the strongest clustering structure and the lowest reconstruction errors, suggesting improved feature representation for automated analysis. Spatial error mapping also revealed that anatomically complex mouthpart structures—such as the mentum, mandibles, ventromental plates, and labrum–epipharynx—consistently present greater reconstruction difficulty.
Overall, this work demonstrates that image modality plays a critical role in optimizing machine learning workflows for forensic entomology and provides a methodological foundation for automated morphological analysis and anomaly detection.
Researcher
Jess Yeap Thong Yi, Ahmad Hadri Jumaat, Raja M. Zuha
Methodology
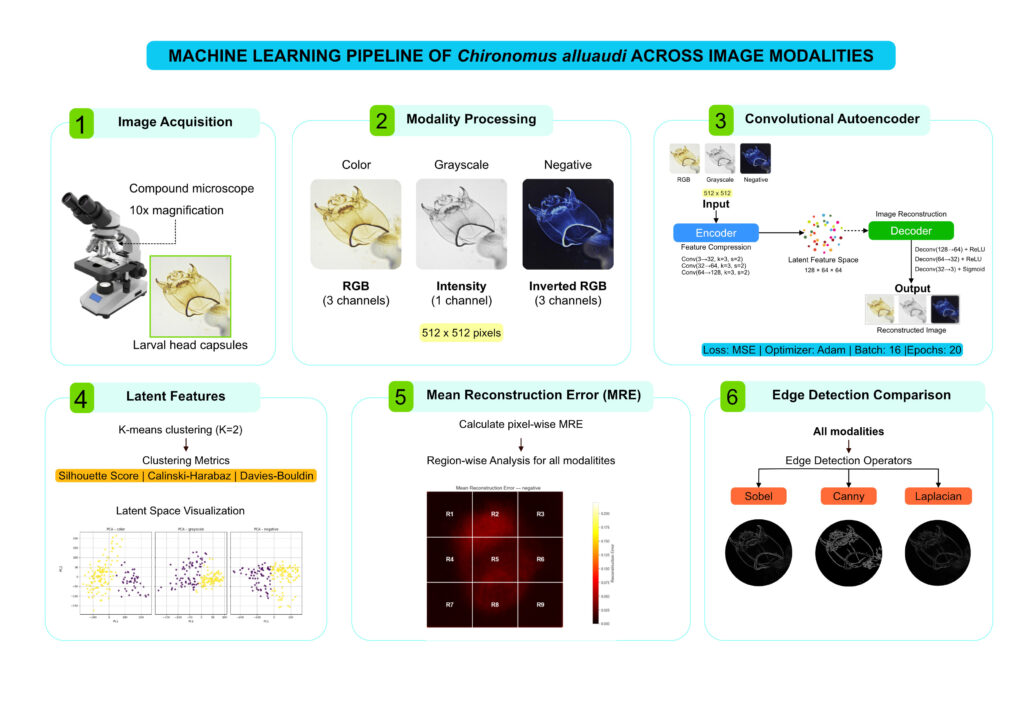
Key Findings
- Image modality strongly affects machine learning performance in the analysis of C. alluaudi larval morphology.
- Negative image modality performed best, producing the strongest feature clustering and the lowest reconstruction errors in the CAE model.
- Complex mouthpart structures (mentum, mandibles, ventromental plates, labrum–epipharynx) consistently showed the highest reconstruction difficulty.
- Edge detection metrics remained consistent across image modalities, indicating that basic structural boundaries were preserved regardless of image type.
Limitations
- Samples are limited to a single age cohort and a single species from laboratory. Morphological features of larval head capsules may vary across age, species and samples from natural environment.
Dataset
Dataset consists of unlabelled processed images of C. alluaudi larval head capsules.
Contact rmzuha@ukm.edu.my to facilitate data sharing.
Repository
This repository is intended for:
– Exploratory research
– Methodological development
– Educational and portfolio demonstration
It is not intended as a production-ready forensic decision system.
License
This project is shared for academic and research purposes.
Please cite appropriately if reused or adapted.
Contact rmzuha@ukm.edu.my for more info.
Acknowledgement
This project was completed using equipment and facility at CSI III Lab, Forensic Science Program, Faculty of Health Sciences, Universiti Kebangsaan Malaysia. This work builds upon standard practices in forensic entomology, environmental monitoring, and open-source scientific computing.